 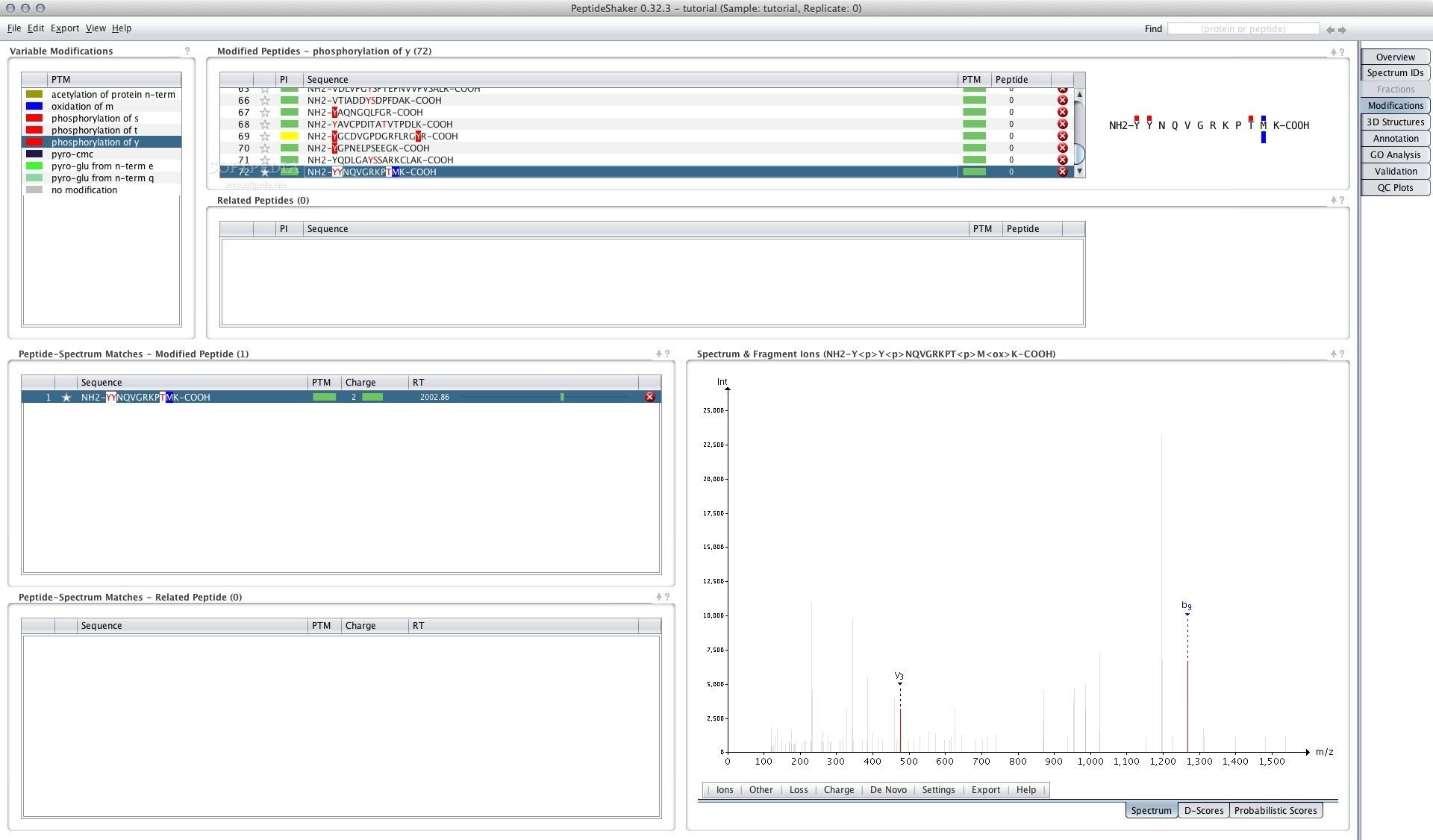
Overview: get a simple yet detailed overview of all the proteins, peptides and PSMs in your dataset.PeptideShaker currently supports nine different analysis tasks: It can be used on local data as well as on data sets deposited to the ProteomeXchange PRIDE repository. PeptideShaker can be used in command line and comes with rich visualization capabilities to navigate the results. PeptideShaker aggregates the results in a single identification set, annotates spectra, computes a consensus score, maps sequences and performs protein inference, scores post-translational modification localization, runs statistical validation, quality control, and annotates the results using multiple sources of information like Gene Ontology, UniProt and Ensembl annotation, and protein structures. PeptideShaker is a search engine independent platform for interpretation of proteomics identification results from multiple search and de novo engines, currently supporting X!Tandem, MS-GF+, MS Amanda, OMSSA, MyriMatch, Comet, Tide, Mascot, Andromeda, MetaMorpheus, Sage, Novor, DirecTag and mzIdentML. (Click on an image to see the full size version) If you use PeptideShaker as part of a publication please include this reference.The reliability of all statistical metrics has been validated in detail using the complex Pyrococcus furiosus standard with. In addition to these identification reliability measures, PeptideShaker also provides confident modification site inference using the latest localization methods (Supplementary Note 1). This filter for specificity and sensitivity includes interactive graphs providing immediate feedback on the values of FDR and FNR for PSMs, peptides and proteins at any chosen threshold. Furthermore, as well as providing false discovery rates (FDRs) at the PSM, peptide and protein levels, PeptideShaker calculates reliable false negative rates (FNRs), providing the user with a novel and highly useful interface to filter results according to an FDR-versus-FNR cost-benefit rationale that has so far been absent from proteomics. PeptideShaker provides statistical confidence estimates for each peptide and protein, taking into account protein inference issues (13). To identify peptides and proteins PeptideShaker uses the target-decoy search strategy (10) to estimate posterior error probabilities and uses these to unify the peptide-to-spectrum match (PSM) lists of different search engines, thus increasing the confidence and sensitivity of hits compared with single-search-engine processing (11,12). Importantly, PeptideShaker can work with the combined output of multiple identification algorithms (Fig. Here we describe PeptideShaker (), a proteomics informatics software that can be used at any stage in the proteomics data cycle for the analysis and interpretation of primary data, enabling data sharing and dissemination and re-analysis of publicly available proteomics data. To maximize the value of public proteomics data, reuse and repurposing must become straightforward, allowing the completion of the proteomics data cycle. There is a pressing need for a user-friendly, open source tool that empowers users to carry out state-of-the-art proteomics data analysis at any stage in the data life cycle (8,9). However, proteomics data processing and (re-)analysis currently remain far from routine practice. Importantly, repository data sharing has enabled the first high-profile studies in which the repurposing of publicly available proteomics data has revealed new biological insights (6,7). Widespread data sharing via publicly accessible repositories, such as PRIDE (1), has now become standard practice, aided by robust user-oriented tools for data submission (2,3) and inspection (4) and bolstered by the advent of the ProteomeXchange initiative (5). Mass spectrometry (MS)-based proteomics is commonly used to identify and quantify the hundreds to thousands of proteins that are present in complex biological samples. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
